Principle of the FTHOM algorithm (H. Hegyi and S. Pongor,
Comput. Applic. Biosci.,9, 371-372, 1992.)
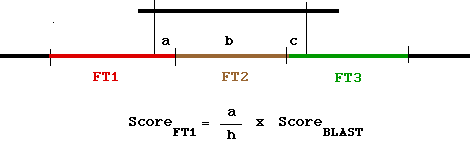
This scoring is repeated for each alignment in the BLAST
output and a sumary list is presented in terms of domain homologies, ranked
by score. The below example is the analysis of the human C1S light chain,
a serin protease which has a propetide removed by proteolysis.
Feature FT Score
name freq
DOMAIN SERINE PROTEASE. 28 6697
DOMAIN CATALYTIC. 1 301
SIMILAR WITH TRYPSIN. 1 269
DOMAIN BETA CHAIN. 1 256
DOMAIN SERINE-PROTEASE. 1 122
DOMAIN SEGMENT A. 1 112
DOMAIN SEGMENT B2. 1 72
DOMAIN SEGMENT B1. 1 62
DOMAIN SEGMENT C. 1 40
PROPEP PROLINE-RICH. 3 27
SITE MAY BE REMOVED BUT IS NOT NECESSARY FOR 1 19
ACTIVATION.
CA_BIND. 1 8
SITE CLEAVAGE SITE (BY FACTOR XA). 3 3
SITE CLEAVAGE (BY FACTOR XIA) (BY SIMILARITY). 2 2
SITE CLEAVAGE (BY FACTOR XIA). 2 2
BINDING SUBSTRATE. 2 2
SITE CLEAVAGE (BY FACTOR XA). 1 1
PROPEP ACTIVATION PEPTIDE. 1 1
SITE CLEAVAGE BY FACTOR XA, FACTOR XIIA, 1 1
SITE THROMBIN ACTIVATION SITE. 1 1
The colors indicate that the different domain names were used in the
early version of Swiss-Prot used for the search. Since 1992, standardization
of domain names started within the SBASE project, and the standard names
greatly simplify the output:
With SBASE 3.0 standard names
SERINE-PROTEASE. 36 7958
PROCESSING-SITE. 14 40
PROPEP PROLINE-RICH. 3 27
CA-BINDING. 1 8
Note that even a short feature, like a proteolitic procesing site of
1-2 amino acids can be noticed with this simple procedure. FTHOM is part
of the domain server (domain@hubi.abc.hu)